-
Notifications
You must be signed in to change notification settings - Fork 0
Expand file tree
/
Copy pathr_environment.Rmd
More file actions
146 lines (109 loc) · 5.16 KB
/
r_environment.Rmd
File metadata and controls
146 lines (109 loc) · 5.16 KB
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
105
106
107
108
109
110
111
112
113
114
115
116
117
118
119
120
121
122
123
124
125
126
127
128
129
130
131
132
133
134
135
136
137
138
139
140
141
142
143
144
145
---
title: "Setting Up R Environment"
output:
html_document:
keep_md: true
theme: cosmo
highlight: tango
toc: true
toc_float: true
toc_depth: 5
df_print: kable
---
```{r setup, include=FALSE}
knitr::opts_chunk$set(echo = TRUE, include = TRUE, message = FALSE, warning = FALSE, tidy = TRUE)
options(digits = 3, scipen = 999)
```
# R Packages
R packages can be thought of as *software* or *applications* that allow you to perform different functions.
## Package installation
Although you have to load your packages into your R environment for every session, you only have to install packages once.
```{r eval=FALSE}
install.packages(c("tidyverse", "here"))
# packages that are not available on CRAN
# install.packages("devtools")
devtools::install_github("hadley/emo")
```
### For PSY2002
These are the packages you would need for PSY2002 lectures.
```{r}
psy2002_packages <- c("plotrix", "car", # lecture 1
"mediation", # lecture 2
"lavaan", "semPlot", # Lecture 3
"lme4", "lmerTest", "emmeans", "e1071", # Lectures 4 & 5
"pscl", "MASS", "boot", # Lecture 6
"cluster", # lecture 9
"psych", # lecture 10
"survival", "km.ci", # lecture 11
"TSA", "TTR", "tseries", "nlme") # lecture 12
```
Let's try installing these packages using `install.packages()` function.
```{r eval=FALSE}
install.packages(psy2002_packages)
```
Once you have them installed, you would only have to "load" them into your environment on an as-needed basis.
## Package loading
You have to load necessary package(s) to your current R environment using `library()` function.
Let's try loading packages we installed for PSY2002.
```{r eval=FALSE}
invisible(lapply(psy2002_packages, # a vector of package names we need for PSY2002
library, # function we want to apply to the vector of multiple packages
character.only = TRUE)) # additional argument for library() function
```
You can check what packages are currently loaded into memory by executing `search()` function.
```{r}
search()
```
Keep track of the package versions with `sessionInfo()`.
```{r}
print(sessionInfo(), locale = FALSE, RNG = FALSE)
```
# R environment
Setting up your R environment from the get-go will prevent any future headaches with respect to importing files and exporting your output.
Your current R session may be opearating at a specific folder, or *working directory* (i.e., WD). Your default working directory may be where your R is installed, like a permanent address. You can change this to a new folder, like a mailing address.
A mailing address is useful for file organization. When you are importing a file (e.g., a dataset) into your R environment, your R will assume the file is located at the working directory. When you are exporting an output or saving a file (e.g., plots, processed datasets), the default location will also be your working directory.
## Working Directory
Let's find where our current R session is operating at.
```{r eval = TRUE}
getwd()
```
We can look at the files available at the current folder by either going to the *Files* tab in the RStudio panel, or run the following codes.
```{r eval=TRUE}
list.files()
dir()
```
## Absolute vs. Relative Path
In our current working directory (i.e., the folder we are operating in), we only have to specify the location **relative** to the working directory (i.e., **relative path**) as opposed to the full location (i.e., **absolute path**).
There is a R package for the relative path, `here`. Install & load the package.
```{r}
# install.packages("here")
library(here)
```
## RStudio Project
To avoid importing and exporting errors, we can create a new project inside the RStudio.
Consider creating a separate project for every new study and for different purposes (e.g., workshops, courses).
You can open the RProject project *project_name.Rproj*, which will oepn a separate RStudio window with the scripts you have been working prior to closing.
For example, you would click on the `r_tutorial.Rproj` when you want to resume working on the project.

### Exercise 1
Let's create a new project for PSY2002.
Step 1: Creating a new project
File → New Project → New Directory
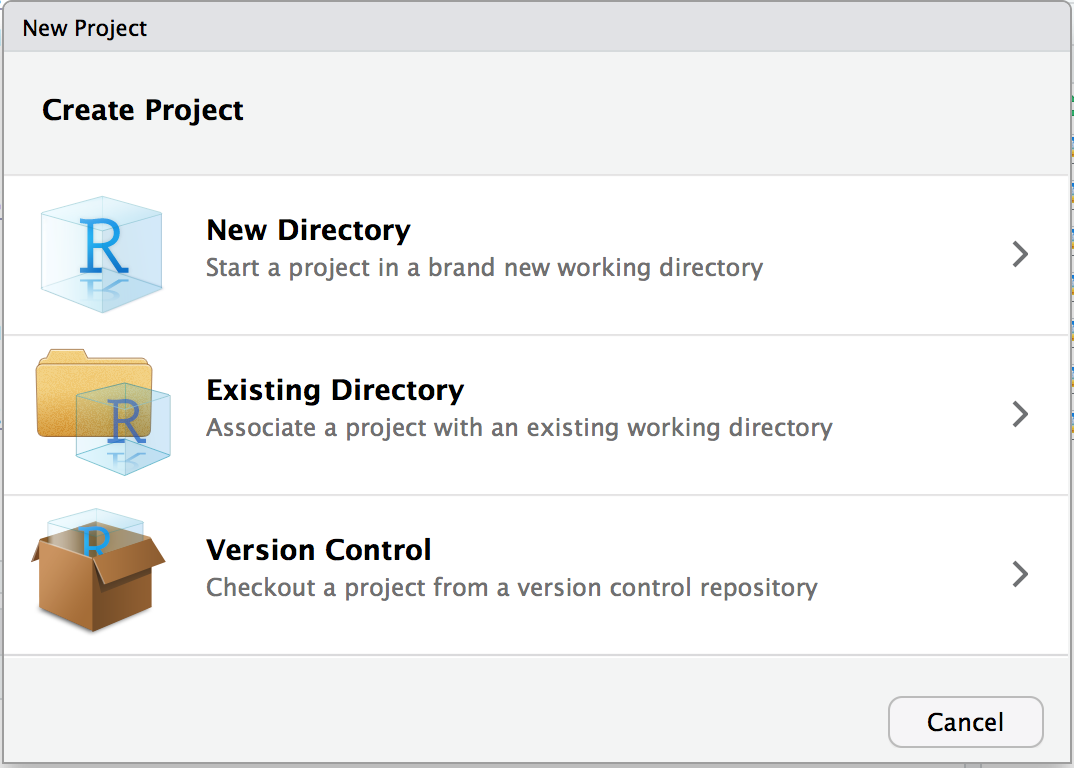
Having a separate RStudio Project will allow you to have a working directory dedicated for a specific projet.
Step 2: Importing a file
Create a new folder called *test_dataset*.
```{r eval=FALSE}
dir.create(here::here("test_dataset"))
```
Then create a copy of the example dataset called **GLMData.csv**, and move it into the new *dataset* folder.
```{r eval=FALSE}
file.copy(from = here::here("dataset", "GLMData.csv"), # the original file & its location
to = here::here("test_dataset", "GLMData.csv")) # the new folder location
```
## Importing files
We can now use a relative path to import files.
Import example dataset "GLMData.csv" located in the dataset folder, and assign an object name (e.g., "df").
```{r eval=FALSE}
df <- read_csv(here::here("test_dataset", "GLMData.csv"))
```